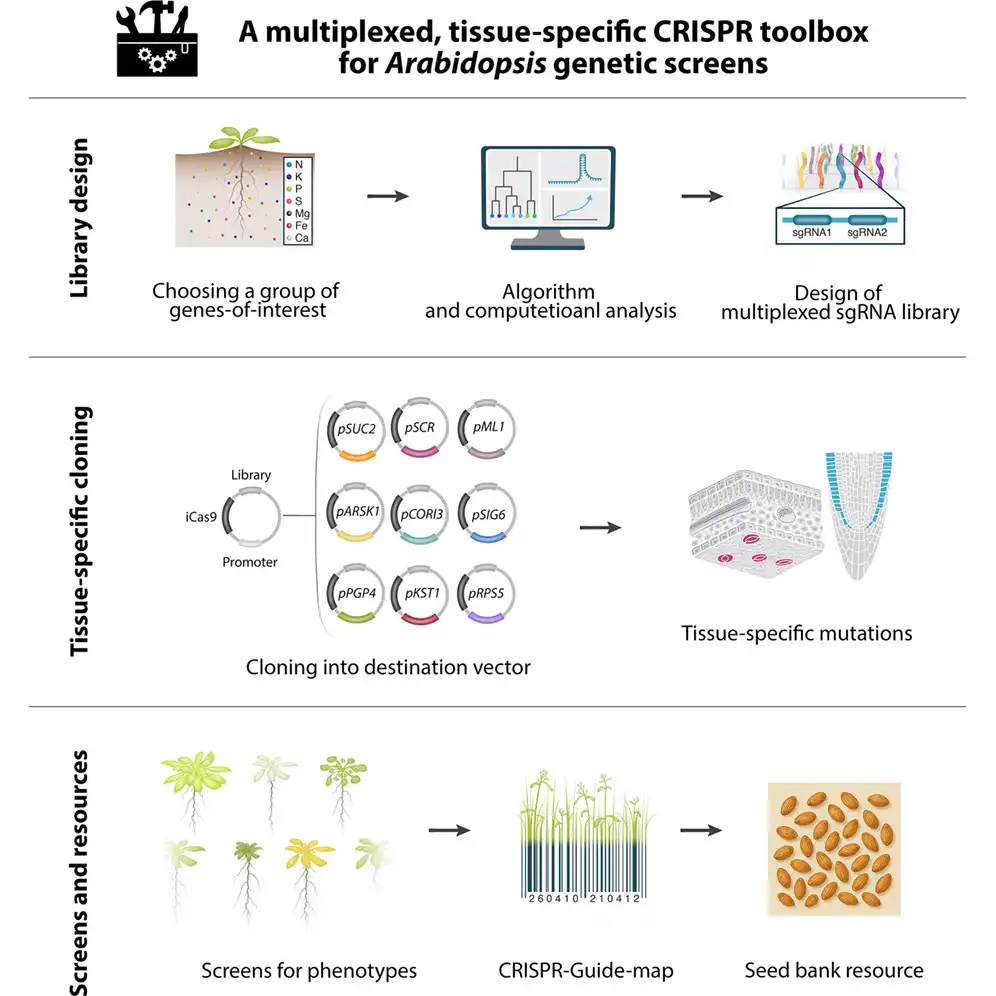
Genome-scale CRISPR libraries are powerful tools for discovering gene function in plants, but traditional designs often edit only one gene at a time, lack spatial control, and struggle to reveal phenotypes when genes act redundantly within families. Here we develop a multiplexed CRISPR library in which each construct carries two sgRNAs that simultaneously target multiple members of a gene family and can be expressed in specific tissues or cell types. A double-barcoding strategy enables efficient tracking of sgRNA combinations at the whole-plant level without sequencing each line individually. Using this platform, we generated over 1,000 Arabidopsis lines targeting 707 transporter genes across 114 gene families involved in nutrient uptake. The multiplexed design improves gene coverage and editing efficiency, revealing phenotypes that are often masked by genetic redundancy and providing a scalable toolbox for spatially precise, multi-gene forward genetic screens in plants.
Anfang M, Yahya RH, Caldararu O, Ben Yaakov S, Landau U, Berman A, Hu Y, Belew ZM, Crocoll C, Xu D, Nour-Eldin HH, Mayrose I, Shani E. Targeting redundant gene families: A multiplexed, tissue-specific CRISPR toolbox for Arabidopsis genetic screens. Cell Rep. 2026 Mar 9;45(3):117055. doi: 10.1016/j.celrep.2026.117055.