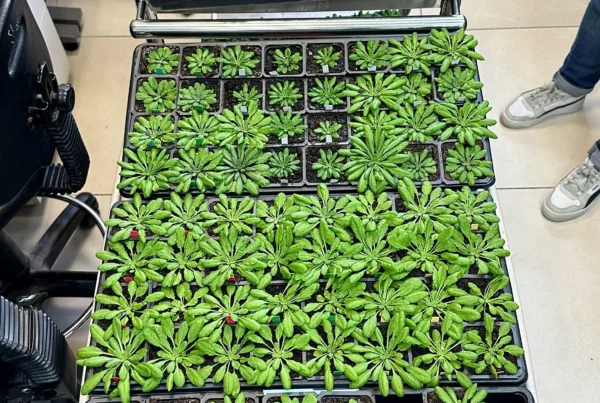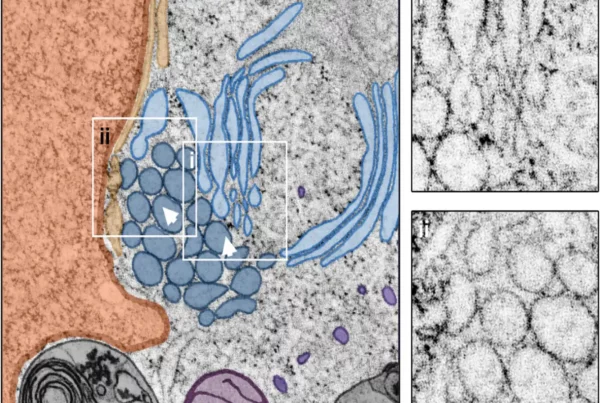
Application of STEM tomography to investigate smooth ER morphology under stress conditions
March 17, 2026
Application of STEM tomography to investigate smooth ER morphology under stress conditions
Heinz, V., Rachel, R., & Ziegler, C. (2025). Application of STEM tomography to investigate smooth ER morphology under stress conditions. Journal of Microscopy, 299(3), 228-241. https://doi.org/10.1111/jmi.70020

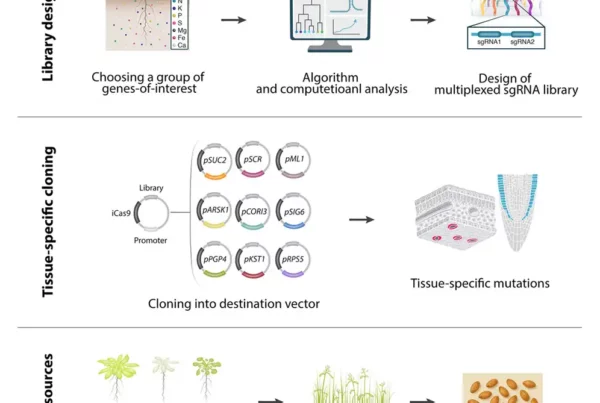
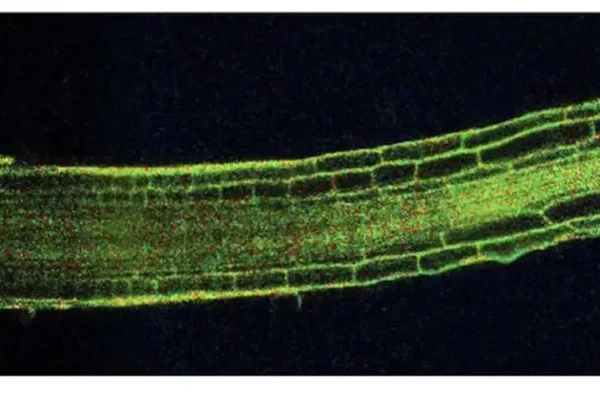


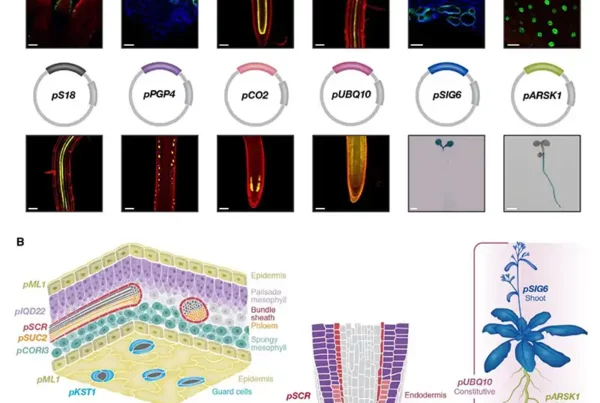
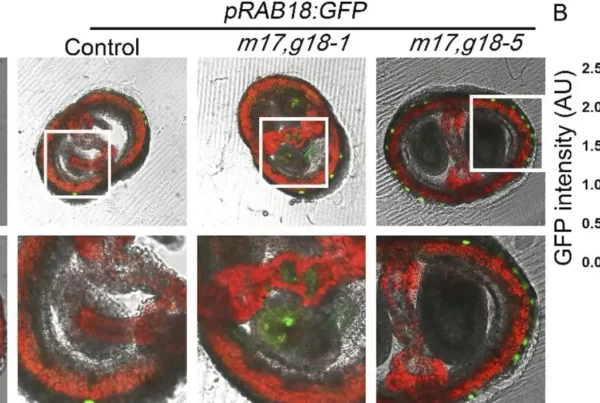
![Image: Description and validation of pMDS plasmid system for dual analysis of transcription and translation in plants. (A) Organization of pMDS1 vector showing reporters for transcription [mTurquoise (mTurQ)], translation (C-terminal mVenus), and a 2A self-cleaving peptide. (B) Organization of pMDS2 vector showing reporters for transcription (mTurQ), translation (N-terminal mVenus), and a 2A self-cleaving peptide. (C) Confocal image of pMDS1_SHRpro:SHR:mVenus:mTurQ showing gene expression (mTurQ) in the stele region and protein (mVenus) translocating to endodermis in the root meristem. (D) Confocal image of pMDS2_VAM3pro:mVenus:VAM3:mTurQ showing subcellular expression (mTurQ) in the nucleus and protein (mVenus) moving to the vacuole in the root epidermis. Red channel shows mCherry expression. nu, nucleus; vac, vacuole; *, endodermis of root meristem. Scale bar, 10 μM. Credit: Science Advances](https://hydrosensing.eu/wp-content/uploads/2026/01/sciadv.adw9153-f1-600x403.webp)